snap-stanford/Biomni

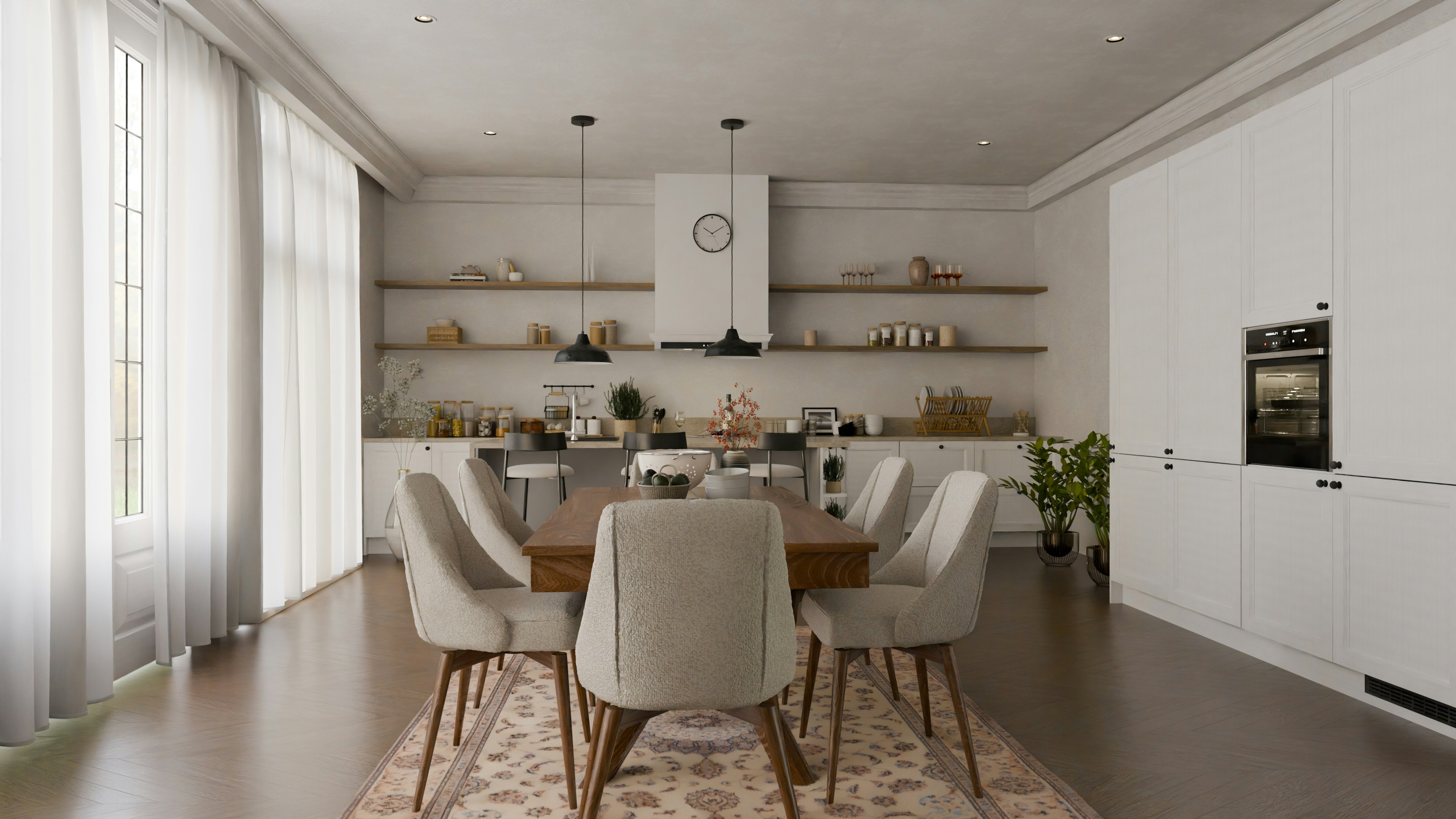
Biomni: A General-Purpose Biomedical AI Agent
Overview
Biomni is a general-purpose biomedical AI agent designed to autonomously execute a wide range of research tasks across diverse biomedical subfields. By integrating cutting-edge large language model (LLM) reasoning with retrieval-augmented planning and code-based execution, Biomni helps scientists dramatically enhance research productivity and generate testable hypotheses.
Quick Start
Installation
Our software environment is massive and we provide a single setup.sh script to setup. Follow this file to setup the env first.
Then activate the environment E1:
|
|
then install the latest biomni package:
|
|
Or install from the github source version.
Lastly, configure your API keys in bash profile ~/.bashrc:
|
|
Basic Usage
Once inside the environment, you can start using Biomni:
|
|
🤝 Contributing to Biomni
Biomni is an open-science initiative that thrives on community contributions. We welcome:
- 🔧 New Tools: Specialized analysis functions and algorithms
- 📊 Datasets: Curated biomedical data and knowledge bases
- 💻 Software: Integration of existing biomedical software packages
- 📋 Benchmarks: Evaluation datasets and performance metrics
- 📚 Misc: Tutorials, examples, and use cases
- 🔧 Update existing tools: many current tools are not optimized - fix and replacements are welcome!
Check out this Contributing Guide on how to contribute to the Biomni ecosystem.
If you have particular tool/database/software in mind that you want to add, you can also submit to this form and the biomni team will implement them.
🔬 Call for Contributors: Help Build Biomni-E2
Biomni-E1 only scratches the surface of what’s possible in the biomedical action space.
Now, we’re building Biomni-E2 — a next-generation environment developed with and for the community.
We believe that by collaboratively defining and curating a shared library of standard biomedical actions, we can accelerate science for everyone.
Join us in shaping the future of biomedical AI agent.
- Contributors with significant impact (e.g., 10+ significant & integrated tool contributions or equivalent) will be invited as co-authors on our upcoming paper in a top-tier journal or conference.
- All contributors will be acknowledged in our publications.
- More contributor perks…
Let’s build it together.
Tutorials and Examples
Biomni 101 - Basic concepts and first steps
More to come!
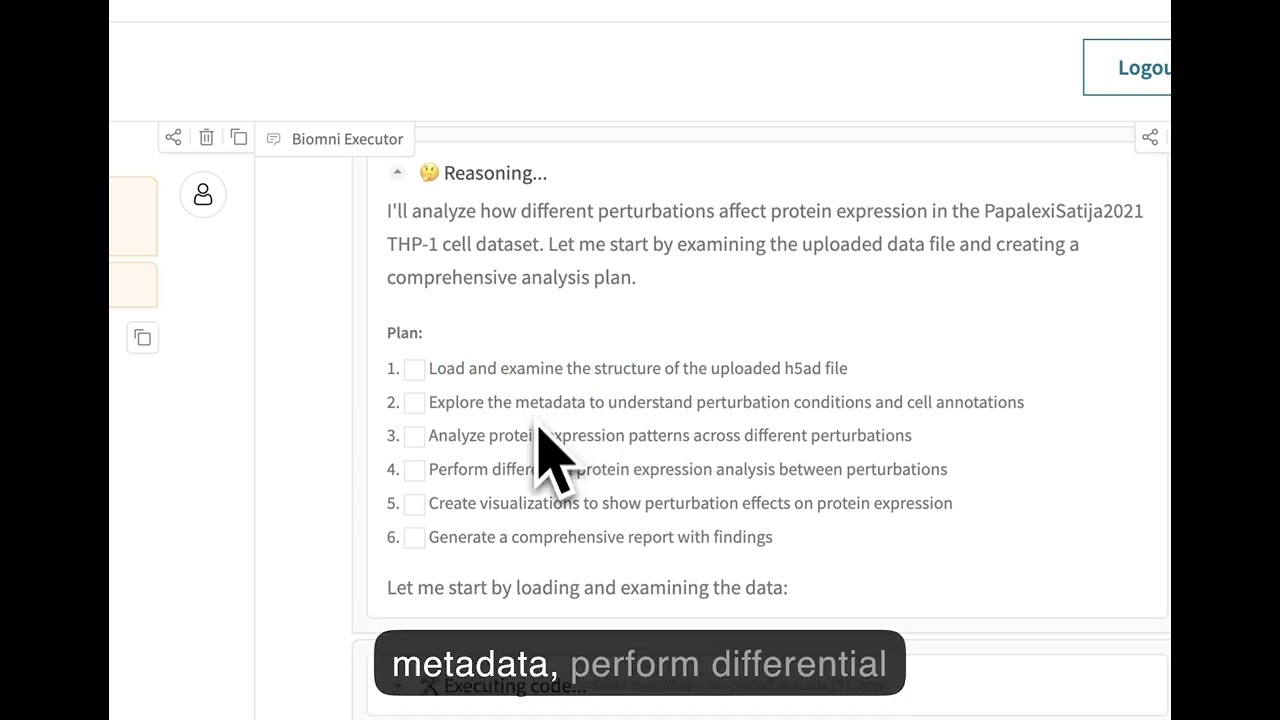
🌐 Web Interface
Experience Biomni through our no-code web interface at biomni.stanford.edu.
Release schedule
- 8 Real-world research task benchmark/leaderboard release
- A tutorial on how to contribute to Biomni
- A tutorial on baseline agents
- Biomni A1+E1 release
Note
- This release was frozen as of April 15 2025, so it differs from the current web platform.
- Biomni itself is Apache 2.0-licensed, but certain integrated tools, databases, or software may carry more restrictive commercial licenses. Review each component carefully before any commercial use.
Cite Us
|
|